This project was funded by the Federal Ministry of Education and Research on the basis of a decision by the German Bundestag.
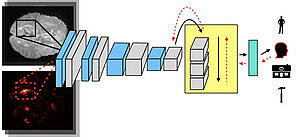
The quantity of data in the health sector (e.g., imagery and ECG time series) is growing exponentially. Machine learning can support the analysis and interpretation of these data so that medical practitioners can create diagnoses more efficiently (see, e.g., MIT Technology Review). In TraMeExCo, the robustness and transparency of diagnostic prediction through machine learning is explored. For two clinical fields (pathology and pain analysis), machine learning methods are tested with three types of data (microscopy images, pain videos, and ECG time series). First, deep learning methods are combined with "white-box" learning methods. Then, clearly defined learned classifiers are used to understand the decisions made by the system. Finally, Bayesian deep learning is used to explore the underlying uncertainties of the system and data. For these applications, two prototype "Transparent Companions for Medical Applications" are developed.
Project Partners
- Fraunhofer IIS
- University of Bamberg
Publications
| [1] | Jeroen Aeles, Fabian Horst, Sebastian Lapuschkin, Lilian Lacourpaille, and François Hug. “Revealing the unique features of each individual’s muscle activation signatures”. In: Journal of the Royal Society Interface 18.174 (Jan. 2021), p. 20200770. DOI: 10.1098/rsif.2020.0770. URL: https://hal.archives-ouvertes.fr/hal-03298503. |
| [2] | Christopher J. Anders, Leander Weber, David Neumann, Wojciech Samek, Klaus-Robert Müller, and Sebastian Lapuschkin. “Finding and removing Clever Hans: Using explanation methods to debug and improve deep models”. In: Information Fusion 77 (2022), pp. 261–295. issn: 1566-2535. DOI: https://doi.org/10.1016/j.inffus.2021.07.015. URL: https://www.sciencedirect.com/science/article/pii/S1566253521001573. |
| [3] | Miriam Hägele, Philipp Seegerer, Sebastian Lapuschkin, Michael Bockmayr, Wojciech Samek, Frederick Klauschen, Klaus-Robert Müller, and Alexander Binder. “Resolving challenges in deep learning-based analyses of histopathological images using explanation methods”. In: Scientific Reports 10.1 (Apr. 2020), p. 6423. ISSN: 2045-2322. DOI: 10.1038/s41598-020-62724-2. URL: https://doi.org/10.1038/s41598-020-62724-2. |
| [4] | Fabian Horst, Sebastian Lapuschkin, Wojciech Samek, Klaus-Robert Müller, and Wolfgang I. Schöllhorn. “Explaining the unique nature of individual gait patterns with deep learning”. In: Scientific Reports 9.1 (Feb. 2019), p. 2391. ISSN: 2045-2322. DOI: 10.1038/s41598- 019- 38748- 8. URL: https://doi.org/10.1038/s41598-019-38748-8. |
| [5] | Maximilian Kohlbrenner, Alexander Bauer, Shinichi Nakajima, Alexander Binder, Wojciech Samek, and Sebastian Lapuschkin. “Towards Best Practice in Explaining Neural Network Decisions with LRP”. In: July 2020, pp. 1–7. DOI: 10.1109/IJCNN48605.2020.9206975. |
| [6] | Grégoire Montavon, Wojciech Samek, and Klaus-Robert Müller. “Methods for interpreting and understanding deep neural networks”. In: Digital Signal Processing 73 (2018), pp. 1–15. ISSN: 1051-2004. DOI: https://doi.org/10.1016/j.dsp.2017.10.011. URL: https://www.sciencedirect.com/science/article/pii/S1051200417302385. |
| [7] | Wojciech Samek, Alexander Binder, Gregoire Montavon, Sebastian Lapuschkin, and Klaus-Robert Müller. “Evaluating the Visualization of What a Deep Neural Network Has Learned”. In: IEEE Transactions on Neural Networks and Learning Systems 28 (Nov. 2017), pp. 2660–2673. DOI: 10.1109/TNNLS.2016.2599820. |
| [8] | Nils Strodthoff and Claas Strodthoff. “Detecting and interpreting myocardial infarction using fully convolutional neural networks”. In: Physiological Measurement 40 (Nov. 2018). DOI: 10.1088/1361-6579/aaf34d. |
| [9] | Jiamei Sun, Sebastian Lapuschkin, Wojciech Samek, and Alexander Binder. “Explain and improve: LRP-inference fine-tuning for image captioning models”. In: Information Fusion 77 (2022), pp. 233–246. ISSN: 1566-2535. DOI: https://doi.org/10.1016/j.inffus.2021.07.008. URL: https://www.sciencedirect.com/science/article/pii/S1566253521001494. |
| [10] | Seul-Ki Yeom, Philipp Seegerer, Sebastian Lapuschkin, Alexander Binder, Simon Wiedemann, Klaus-Robert Müller, and Wojciech Samek. “Pruning by explaining: A novel criterion for deep neural network pruning”. In: Pattern Recognition 115 (2021), p. 107899. ISSN: 0031-3203. DOI: https://doi.org/10.1016/j.patcog.2021.107899. URL: https://www.sciencedirect.com/science/article/pii/S0031320321000868. |